Computational prediction of potential inhibitors for SARS-COV-2 main protease based on machine learning, docking, MM-PBSA calculations, and metadynamics | PLOS ONE

Potential repurposing of four FDA approved compounds with antiplasmodial activity identified through proteome scale computational drug discovery and in vitro assay | Scientific Reports
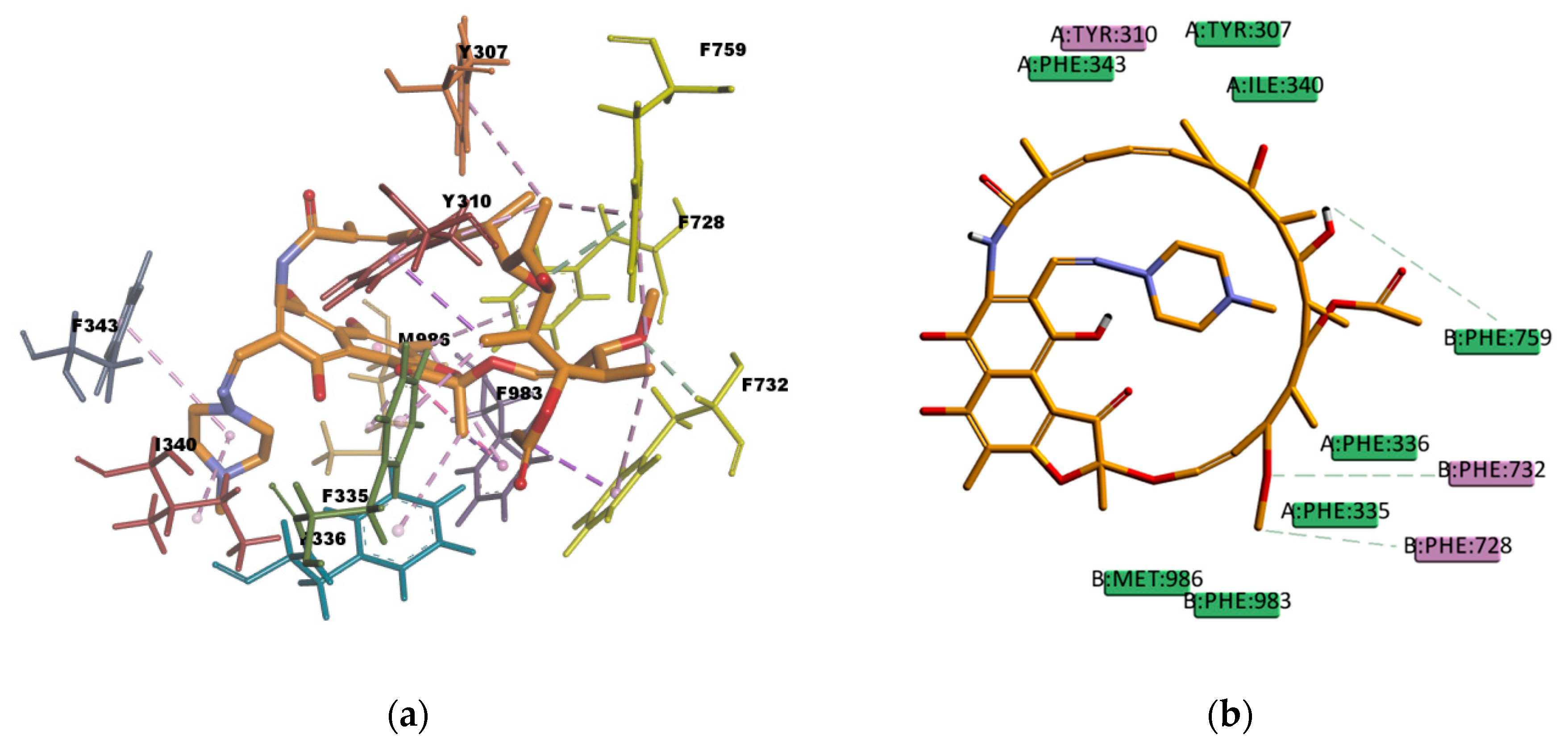
![Virtual screening, optimization and molecular dynamics analyses highlighting a pyrrolo[1,2-a]quinazoline derivative as a potential inhibitor of DNA gyrase B of Mycobacterium tuberculosis | Scientific Reports Virtual screening, optimization and molecular dynamics analyses highlighting a pyrrolo[1,2-a]quinazoline derivative as a potential inhibitor of DNA gyrase B of Mycobacterium tuberculosis | Scientific Reports](https://media.springernature.com/full/springer-static/image/art%3A10.1038%2Fs41598-022-08359-x/MediaObjects/41598_2022_8359_Fig1_HTML.png)
Virtual screening, optimization and molecular dynamics analyses highlighting a pyrrolo[1,2-a]quinazoline derivative as a potential inhibitor of DNA gyrase B of Mycobacterium tuberculosis | Scientific Reports

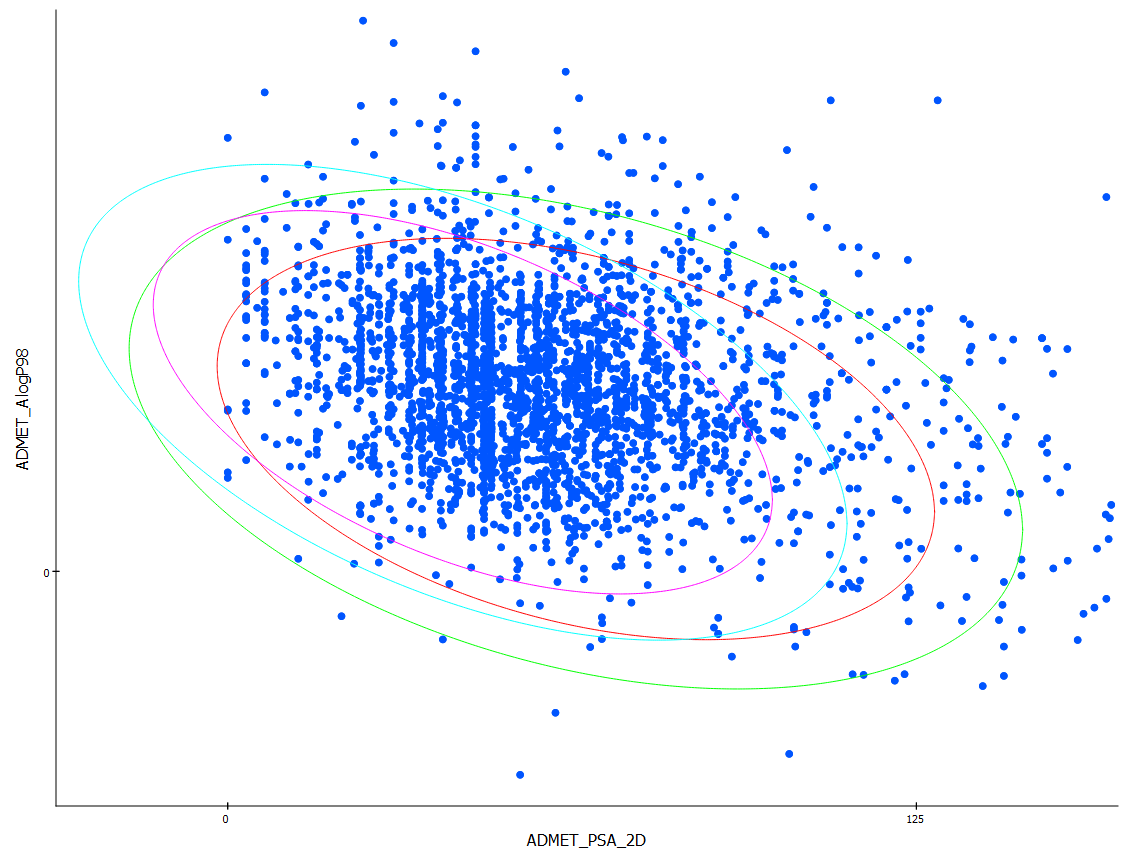
Structure based pharmacophore modeling, virtual screening, molecular docking and ADMET approaches for identification of natural anti-cancer agents targeting XIAP protein | Scientific Reports
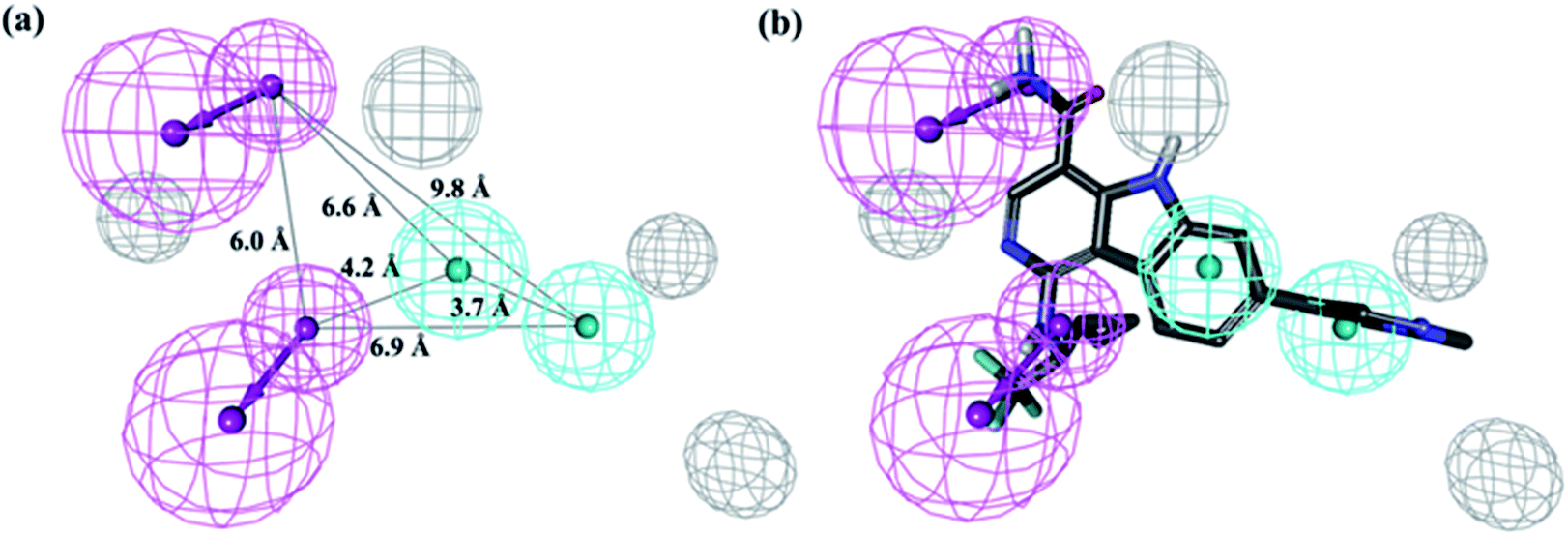
Integration of pharmacophore mapping and molecular docking in sequential virtual screening: towards the discovery of novel JAK2 inhibitors - RSC Advances (RSC Publishing) DOI:10.1039/C6RA24959K
![Computer-aided molecular modeling studies of some 2, 3-dihydro-[1, 4] dioxino [2, 3-f] quinazoline derivatives as EGFRWT inhibitors | Beni-Suef University Journal of Basic and Applied Sciences | Full Text Computer-aided molecular modeling studies of some 2, 3-dihydro-[1, 4] dioxino [2, 3-f] quinazoline derivatives as EGFRWT inhibitors | Beni-Suef University Journal of Basic and Applied Sciences | Full Text](https://media.springernature.com/lw685/springer-static/image/art%3A10.1186%2Fs43088-020-00047-x/MediaObjects/43088_2020_47_Fig4_HTML.png)
Computer-aided molecular modeling studies of some 2, 3-dihydro-[1, 4] dioxino [2, 3-f] quinazoline derivatives as EGFRWT inhibitors | Beni-Suef University Journal of Basic and Applied Sciences | Full Text

Ligand-based Pharmacophore Modeling, Virtual Screening and Molecular Docking Studies for Discovery of Potential Topoisomerase I Inhibitors - ScienceDirect