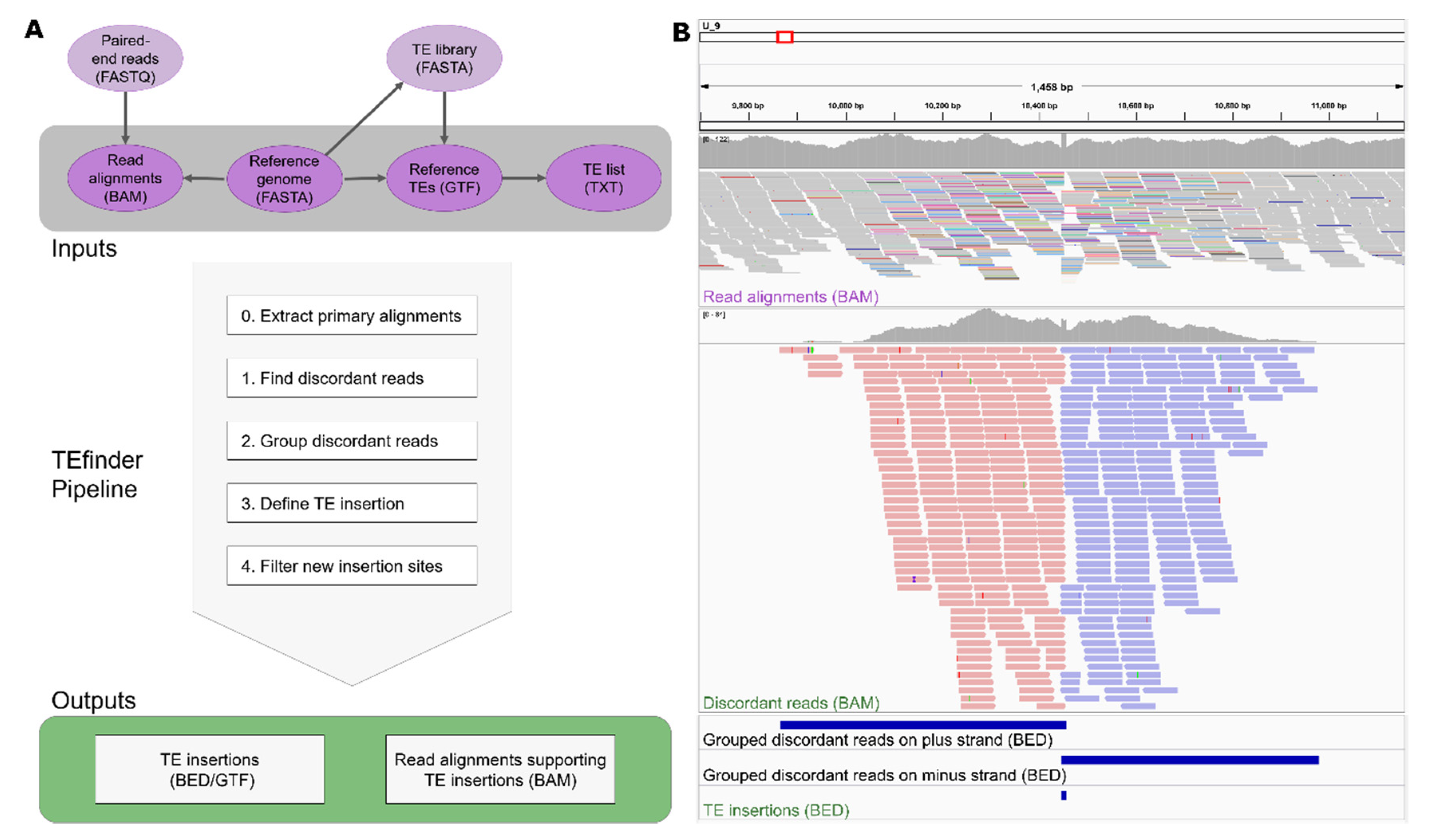
Genes | Free Full-Text | TEfinder: A Bioinformatics Pipeline for Detecting New Transposable Element Insertion Events in Next-Generation Sequencing Data

Validation of a Customized Bioinformatics Pipeline for a Clinical Next-Generation Sequencing Test Targeting Solid Tumor–Associated Variants - ScienceDirect

The use of technical replication for detection of low-level somatic mutations in next-generation sequencing | Nature Communications

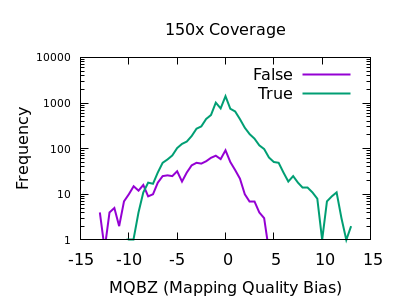
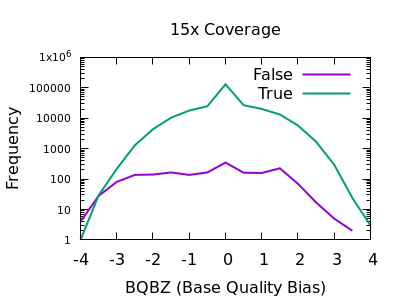
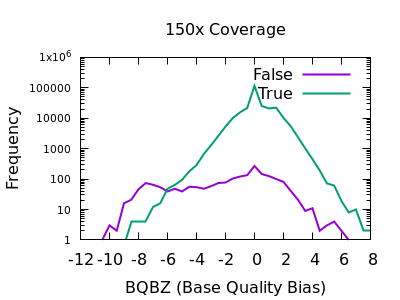
Allele and strand bias for SNVs. This figure shows read distribution of... | Download Scientific Diagram

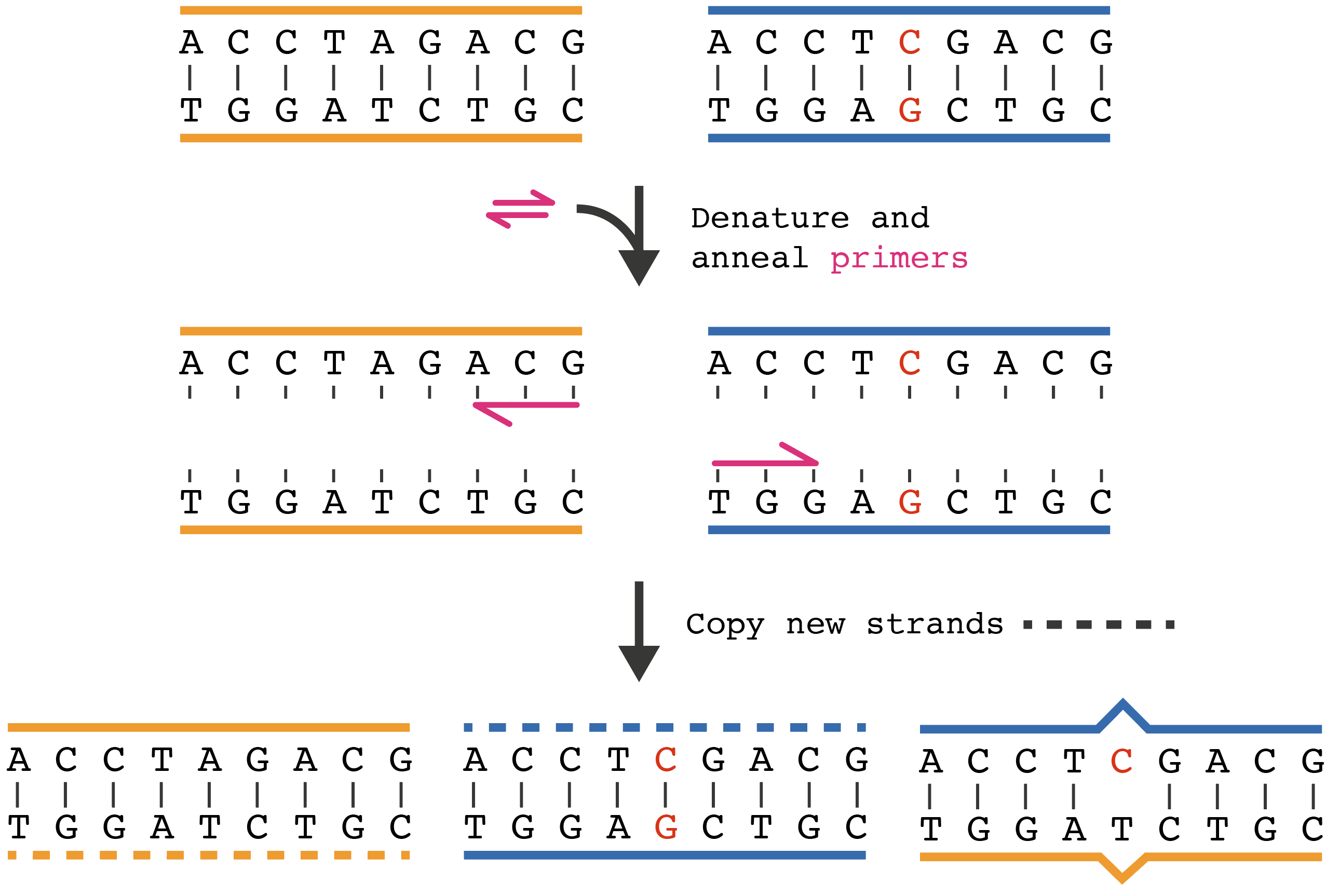
Common artifacts in NGS alignments that gave rise to a false-positive... | Download Scientific Diagram

Validation of a Customized Bioinformatics Pipeline for a Clinical Next-Generation Sequencing Test Targeting Solid Tumor–Associated Variants - ScienceDirect

HIV-1 Drug Resistance Assay Using Ion Torrent Next Generation Sequencing and On-Instrument End-to-End Analysis Software | Journal of Clinical Microbiology