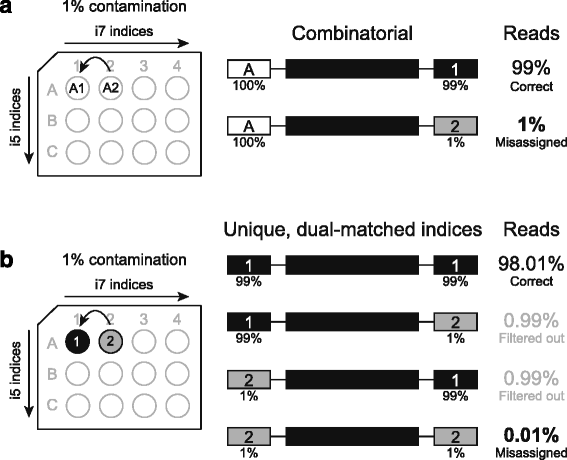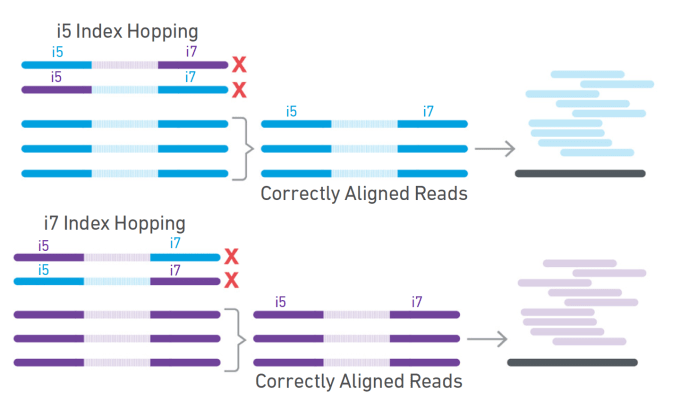
Unique, dual-indexed sequencing adapters with UMIs effectively eliminate index cross-talk and significantly improve sensitivity of massively parallel sequencing | BMC Genomics | Full Text
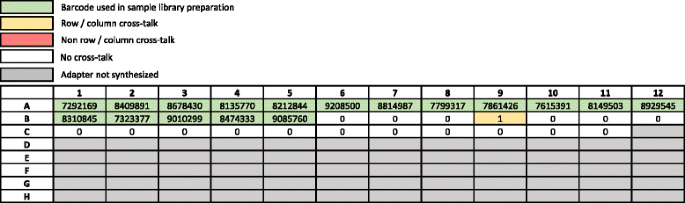
Unique, dual-indexed sequencing adapters with UMIs effectively eliminate index cross-talk and significantly improve sensitivity of massively parallel sequencing | BMC Genomics | Full Text

PDF) Unique, dual-indexed sequencing adapters with UMIs effectively eliminate index cross-talk and significantly improve sensitivity of massively parallel sequencing
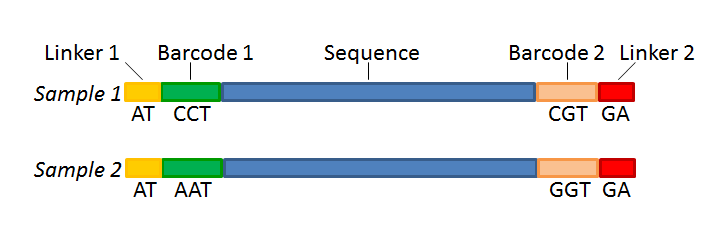
Not All Sequence Tags Are Created Equal: Designing and Validating Sequence Identification Tags Robust to Indels | PLOS ONE
Not All Sequence Tags Are Created Equal: Designing and Validating Sequence Identification Tags Robust to Indels | PLOS ONE
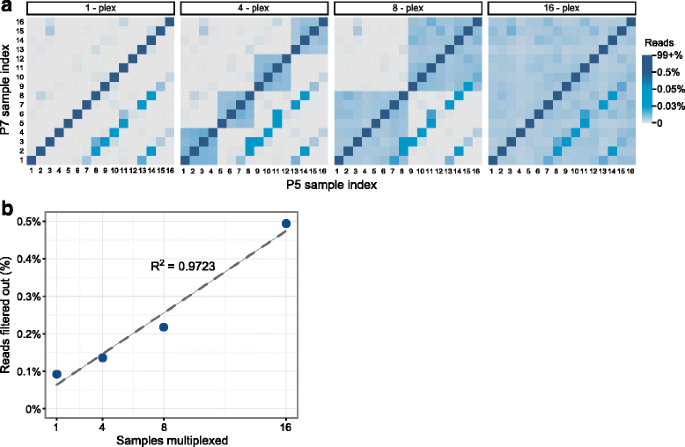
Unique, dual-indexed sequencing adapters with UMIs effectively eliminate index cross-talk and significantly improve sensitivity of massively parallel sequencing | BMC Genomics | Full Text
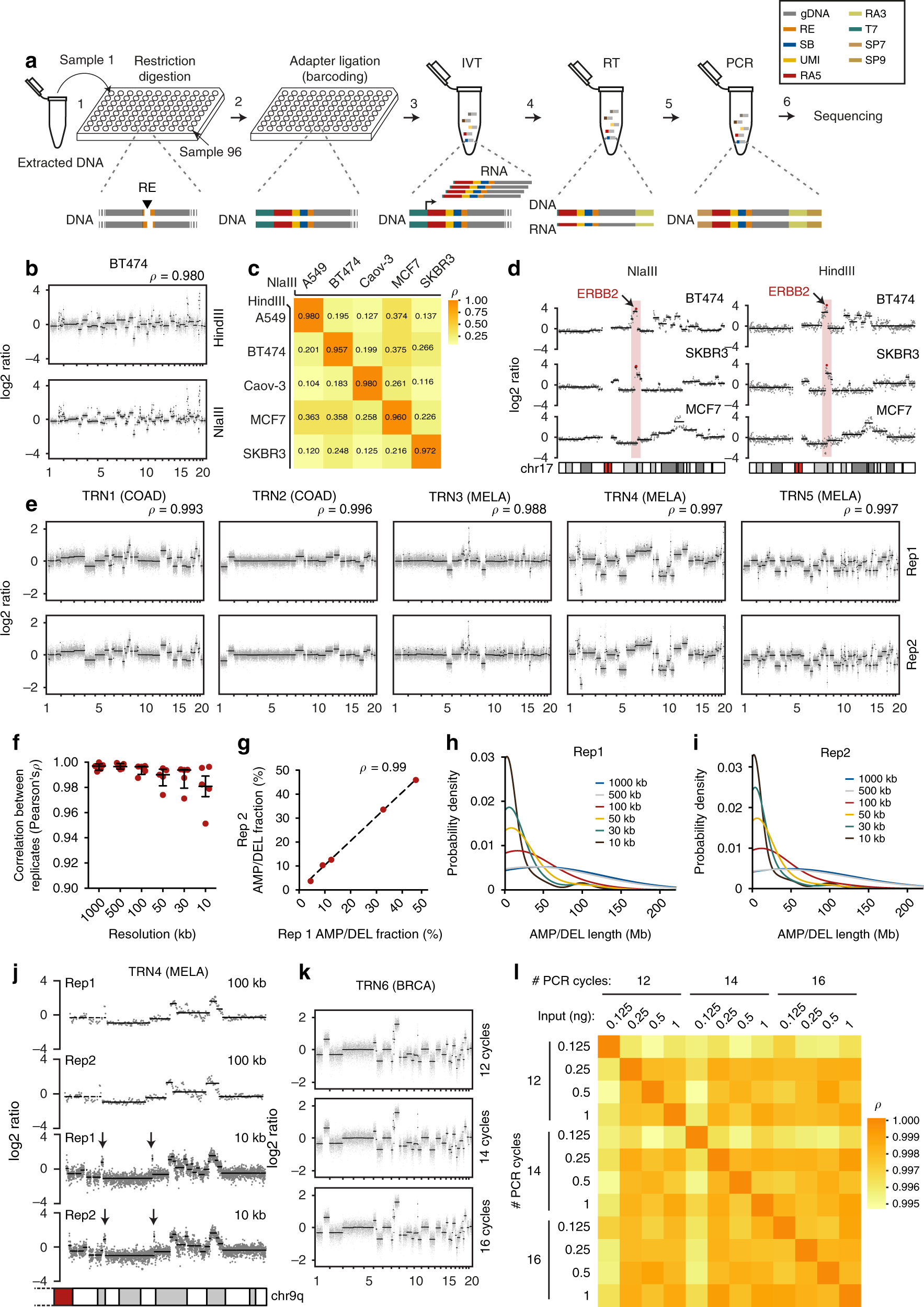
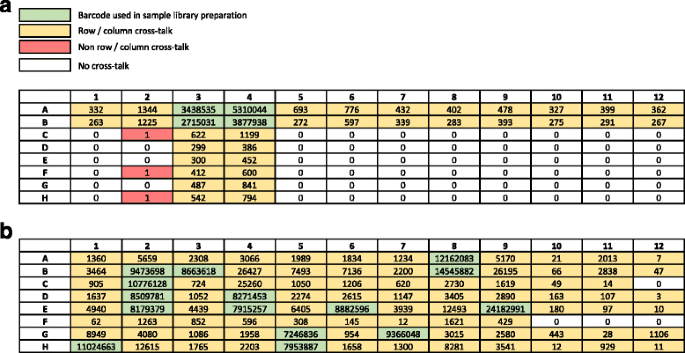
CUTseq is a versatile method for preparing multiplexed DNA sequencing libraries from low-input samples | Nature Communications
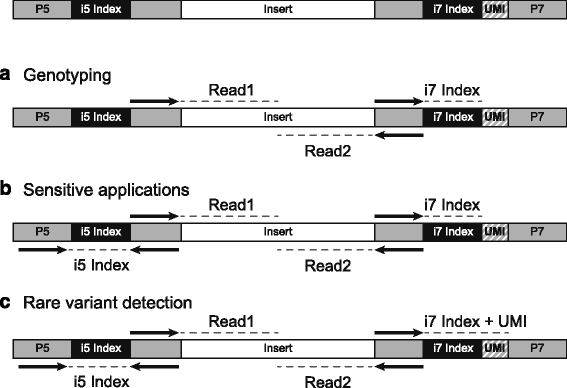
Unique, dual-indexed sequencing adapters with UMIs effectively eliminate index cross-talk and significantly improve sensitivity of massively parallel sequencing | BMC Genomics | Full Text

Multiplexed Competition in a Synthetic Squid Light Organ Microbiome Using Barcode-Tagged Gene Deletions | mSystems